Laboratory of Photosynthesis
Josef Komenda`s group
Biogenesis of Photosystem II
Research Projects
Phosphonate Biosynthesis and Degradation – A Novel Mitochondrial Pathway Involved in Global Phosphorus Turnover?
Founded by the Czech Science Foundation - Junior Star Project, 2021-2025.
Principal investigator: Jan Janouškovec, Ph.D.
Phosphorus is a core, limiting macronutrient, but its global turnover is still little understood. For example, one quarter of all phosphorus in the ocean occurs as phosphonates, whose metabolism has only been thoroughly studied in bacteria. We identified unexpectedly widespread phosphonate enzymes in eukaryotic genomes and predict that they comprise a novel mitochondrial pathway. To test this, we will computationally map phosphonate enzyme distribution, expression, and cellular localization in all eukaryotes, then localize them subcellularly in two distantly related protists, Perkinsus and Capsaspora. We will next use labeled precursors to track phosphonate metabolism in both species, and measure the ability of a wide range of algae to utilize phosphonate substrates for growth. We will also reconstruct the evolutionary history of all phosphonate enzymes and how they relate to eukaryogenesis. Our findings will illuminate mitochondrial metabolism, eukaryotic evolution, and phosphorus turnover in cells and ecosystems, and identify new enzymes of potential commercial interest.
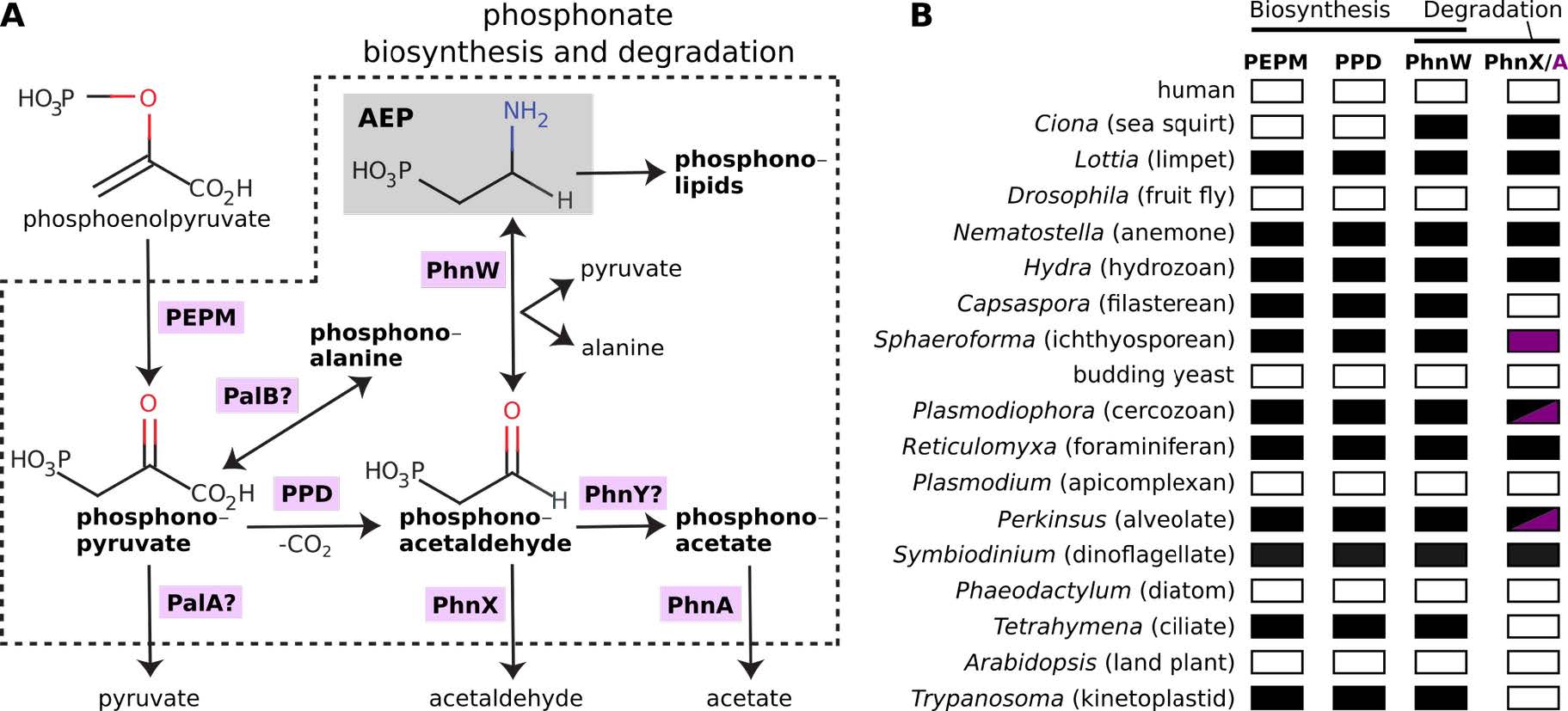
A) Overview of phosphonate metabolism. Enzymes (blue background) are abbreviated as in the text. Question marks denotes enzymes that have not yet been searched in eukaryotes. The most common phosphonate in nature, AEP (2-aminoethylphosphonate), is highlighted in grey. B) Preliminary inquiry into the distribution of the five main genes for phosphonate biosynthesis and degradation in 18 eukaryotic genomes. Filled boxes denote gene presence (purple is used for PhnA) and empty boxes denote absence.
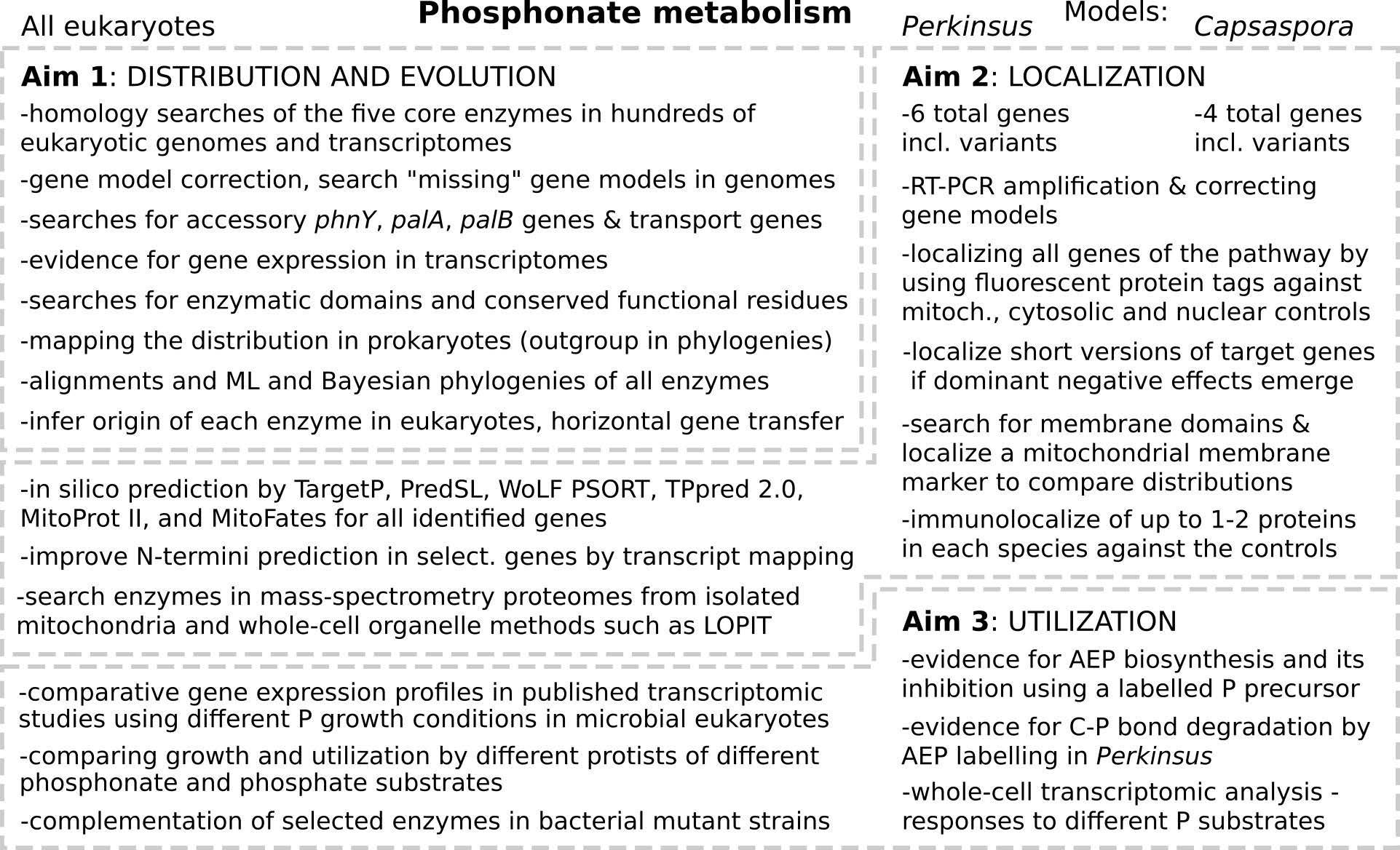

Overview of the three research aims of the project.